Download PDF
Sometimes you have too much information, as a molecular investigation of families with inherited retinal disease demonstrates.1 This retrospective analysis identified disease-causing genotypes in 760 of the 1,000 consecutive families treated by a single clinician, a sensitivity well beyond the 5% chance of a molecular diagnosis when the field was young.
Advances in genome sequencing pose a new set of challenges to clinicians, said Edwin M. Stone, MD, PhD, at the University of Iowa Stephen A. Wynn Institute for Vision Research in Iowa City. There’s a common misconception that as diagnostic modalities evolve, the need for good clinical skills diminishes, he said. In fact, the opposite is true.
“Good clinical skills are more important than ever for arriving at a correct molecular diagnosis,” Dr. Stone said. “Because of the large amount of noise in each person’s genome, one needs a quite focused clinical hypothesis to obtain a statistically significant result from a broad genetic test like whole exome sequencing.”
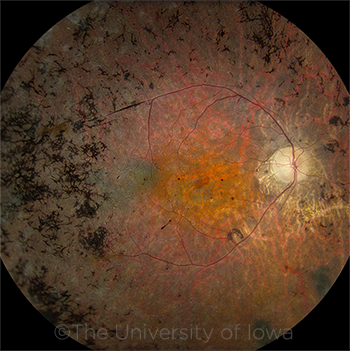 |
RETINITIS PIGMENTOSA. A 64-year-old male with USH2A-associated RP. Note the characteristic lacy black intraretinal pigment, narrowed arterioles, and slightly pale optic disc.
|
Study specifics. While more than 300 genes are currently known to cause inherited retinal disorders, only 104 genes were observed in this study population. Of those, 13 genes were responsible for nearly half of disease in all families (497 families).
Want to hear more from Dr. Stone?
Attend “Genetic Disorders in Pediatric Ophthalmology” at AAO 2017!
Register now
The researchers compared 2 testing strategies: 1) Whole exome sequencing, a single test that can examine hundreds of retinal disease–causing genes at once; and 2) tiered testing, which relies on a good clinical diagnosis to direct the molecular testing. The latter approach narrows the number of genes under consideration, thereby increasing the statistical significance of the results.
Accuracy. The results showed that the tiered testing strategy was 6.1% more sensitive than whole exome sequencing alone. It also resulted in a much lower false genotype rate (FGR). “All clinical tests have some false positives, and molecular tests are no exception,” said Dr. Stone. The study showed the likelihood of observing a plausible disease-causing “result” purely by chance to be 128%, if one tested 300 genes at a time, as current retinal disease sequencing panels can do. Using a tiered approach, the FGR fell below 5%.
Cost. The study also found that a tiered approach was 17.7% less expensive than whole exome sequencing. The latter cost $1,200 per patient. In contrast, customized testing based on clinical findings cost $990 on average.
However, cost is not the reason Dr. Stone advocates a refined testing strategy. “The main value of the pretest hypothesis is to reduce the number of false positives and thereby increase the statistical significance of the results.”
Should you order testing? Retina specialists could use the classification system reported in this study directly, said Dr. Stone. Other clinicians will have to customize it a bit for their practices. All physicians should try to establish the narrowest possible clinical diagnosis before performing a molecular test, he said.
—Miriam Karmel
___________________________
1 Stone EM et al. Ophthalmology. 2017;124(9):1314-1331.
___________________________
Relevant financial disclosures—Dr. Stone: None.
For full disclosures and disclosure key, see below.
More from this month’s News in Review